gemclus.DiscriminativeModel¶
- class gemclus.DiscriminativeModel(n_clusters=3, gemini='mmd_ova', max_iter=1000, learning_rate=0.001, solver='adam', batch_size=None, verbose=False, random_state=None)[source]¶
- This is the BaseGEMINI to derive to create a GEMINI MLP or linear clustering model.
When deriving this class, there are a couple methods to override depending on the design of a discriminative model \(p(y|x)\) or a specific GEMINI.
- Parameters:
- n_clustersint, default=3
The maximum number of clusters to form as well as the number of output neurons in the neural network.
- gemini: str, GEMINI instance or None, default=”mmd_ova”
GEMINI objective used to train this discriminative model. Can be “mmd_ova”, “mmd_ovo”, “wasserstein_ova”, “wasserstein_ovo”, “mi” or other GEMINI available in gemclus.gemini.AVAILABLE_GEMINI. Default GEMINIs involve the Euclidean metric or linear kernel. To incorporate custom metrics, a GEMINI can also be passed as an instance. If set to None, the GEMINI will be MMD OvA with linear kernel.
- max_iter: int, default=1000
Maximum number of epochs to perform gradient descent in a single run.
- learning_rate: float, default=1e-3
Initial learning rate used. It controls the step-size in updating the weights.
- solver: {‘sgd’,’adam’}, default=’adam’
The solver for weight optimisation.
‘sgd’ refers to stochastic gradient descent.
‘adam’ refers to a stochastic gradient-based optimiser proposed by Kingma, Diederik and Jimmy Ba.
- batch_size: int, default=None
The size of batches during gradient descent training. If set to None, the whole data will be considered.
- verbose: bool, default=False
Whether to print progress messages to stdout
- random_state: int, RandomState instance, default=None
Determines random number generation for weights and bias initialisation. Pass an int for reproducible results across multiple function calls.
- Attributes:
- optimiser_: `AdamOptimizer` or `SGDOptimizer`
The optimisation algorithm used for training depending on the chosen solver parameter.
- labels_: ndarray of shape (n_samples)
The labels that were assigned to the samples passed to the
fit()method.- n_iter_: int
The number of iterations that the model took for converging.
- __init__(n_clusters=3, gemini='mmd_ova', max_iter=1000, learning_rate=0.001, solver='adam', batch_size=None, verbose=False, random_state=None)[source]¶
- fit(X, y=None)[source]¶
Compute GEMINI clustering.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Training instances to cluster.
- yndarray of shape (n_samples, n_samples), default=None
Use this parameter to give a precomputed affinity metric if the option “precomputed” was passed during construction. Otherwise, it is not used and present here for API consistency by convention.
- Returns:
- selfobject
Fitted estimator.
- fit_predict(X, y=None)[source]¶
Compute GEMINI clustering and returns the predicted clusters.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Training instances to cluster.
- yndarray of shape (n_samples, n_samples), default=None
Use this parameter to give a precomputed affinity metric if the option “precomputed” was passed during construction. Otherwise, it is not used and present here for API consistency by convention.
- Returns:
- y_predndarray of shape (n_samples,)
Vector containing the cluster label for each sample.
- get_gemini()[source]¶
Initialise a
gemclus.GEMINIinstance that will be used to train the model.- Returns:
- gemini: a GEMINI instance
- get_metadata_routing()¶
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]¶
Return the cluster membership of samples. This can only be called after the model was fit to some data.
- Parameters:
- X{array-like, sparse matrix}, shape (n_samples, n_features)
The input samples.
- Returns:
- yndarray of shape (n_samples,)
The label for each sample is the label of the closest sample seen during fit.
- predict_proba(X)[source]¶
Probability estimates that are the output of the neural network p(y|x). The returned estimates for all classes are ordered by the label of classes.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Vector to be scored, where n_samples is the number of samples and n_features is the number of features.
- Returns:
- Tarray-like of shape (n_samples, n_clusters)
Returns the probability of the sample for each cluster in the model.
- score(X, y=None)[source]¶
Return the value of the GEMINI evaluated on the given test data.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Test samples.
- yndarray of shape (n_samples, n_samples), default=None
Use this parameter to give a precomputed affinity metric if the option “precomputed” was passed during construction. Otherwise, it is not used and present here for API consistency by convention.
- Returns:
- scorefloat
GEMINI evaluated on the output of
self.predict(X).
- set_params(**params)¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Examples using gemclus.DiscriminativeModel¶
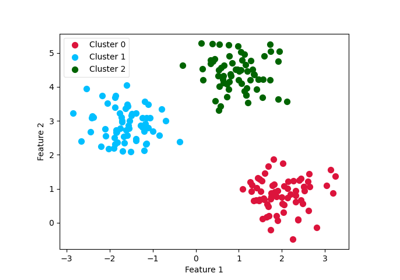
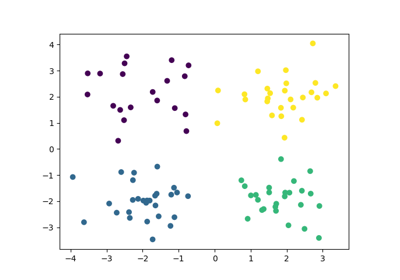
An introducing example to clustering with an MLP and the MMD GEMINI
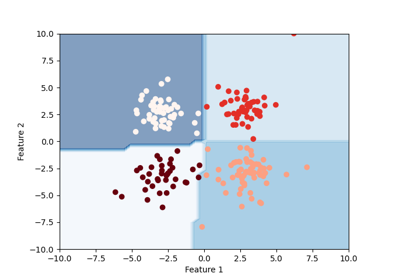
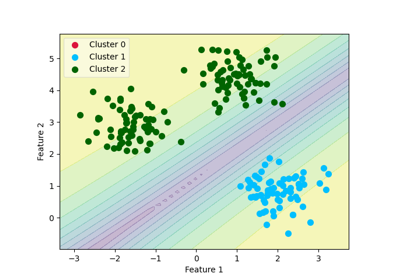
Example of decision boundary map for a mixture of Gaussian and low-degree Student distributions
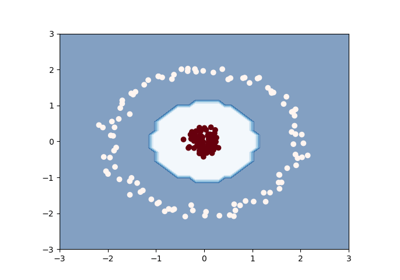
Clustering with the squared-loss mutual information

Drawing a decision boundary between two interlacing moons

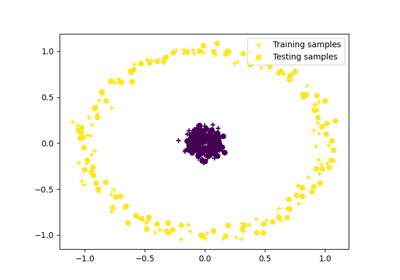
Comparative clustering of circles dataset with kernel change
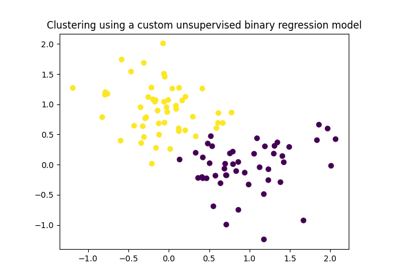
Extending GemClus to build your own discriminative clustering model

Feature selection using the Sparse MMD OvO (Logistic regression)
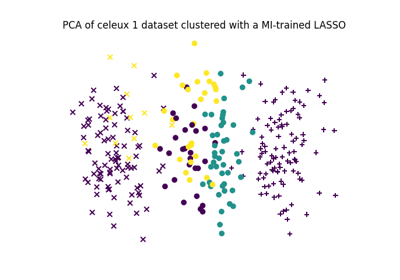
Feature selection using the Sparse Linear MI (Logistic regression)
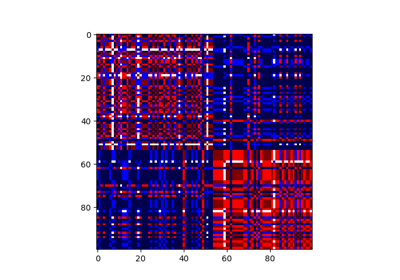
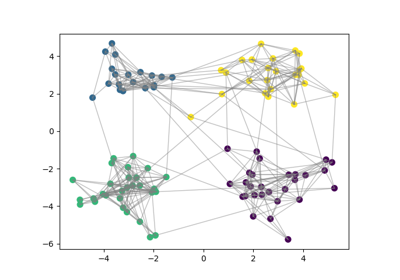
Consensus clustering with linking constraints on sample pairs

Building a differentiable unsupervised tree: DOUGLAS